Making the Logo
Having some funky ellipses in a simplex inspired some interest when I put the logo together for pyrolite, so I put together a cleaned-up example of how you can create these kinds of plots for your own data. These examples illustrate different methods to show distribution of (homogeneous, or near so) compositional data for exploratory analysis.
import matplotlib
import matplotlib.cm
import matplotlib.colors
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pyrolite.plot
from pyrolite.comp.codata import *
from pyrolite.util.plot.helpers import plot_pca_vectors, plot_stdev_ellipses
from pyrolite.util.skl.transform import ALRTransform, ILRTransform
from pyrolite.util.synthetic import random_composition
np.random.seed(82)
# ignore sphinx_gallery warnings
import warnings
warnings.filterwarnings("ignore", "Matplotlib is currently using agg")
First we choose some colors, create some log-distributed synthetic data. Here I’ve
generated a synthetic dataset with four samples having means equidistant from the
log-space centre and with varying covariance. This should illustrate the spatial
warping of the simplex nicely. Additionally, I chose a log-transform here to go
from and to compositional space (ILRTransform, which uses
the isometric log-ratio function ilr()). Choosing
another transform will change the distortion observed in the simplex slightly.
This synthetic dataset is added into a DataFrame for convenient access
to plotting functions via the pandas API defined in pyrolite.plot.pyroplot.
d = 1.0 # distance from centre
sig = 0.1 # scale for variance
# means for logspace (D=2)
means = np.array(np.meshgrid([-1, 1], [-1, 1])).T.reshape(-1, 2) * d
# means = np.array([(-d, -d), (d, -d), (-d, d), (d, d)])
covs = ( # covariance for logspace (D=2)
np.array(
[
[[1, 0], [0, 1]],
[[0.5, 0.15], [0.15, 0.5]],
[[1.5, -1], [-1, 1.5]],
[[1.2, -0.6], [-0.6, 1.2]],
]
)
* sig
)
means = ILRTransform().inverse_transform(means) # compositional means (D=3)
size = 2000 # logo @ 10000
pts = [random_composition(mean=M, cov=C, size=size) for M, C in zip(means, covs)]
T = ILRTransform()
to_log = T.transform
from_log = T.inverse_transform
df = pd.DataFrame(np.vstack(pts))
df.columns = ["SiO2", "MgO", "FeO"]
df["Sample"] = np.repeat(np.arange(df.columns.size + 1), size).flatten()
chem = ["MgO", "SiO2", "FeO"]
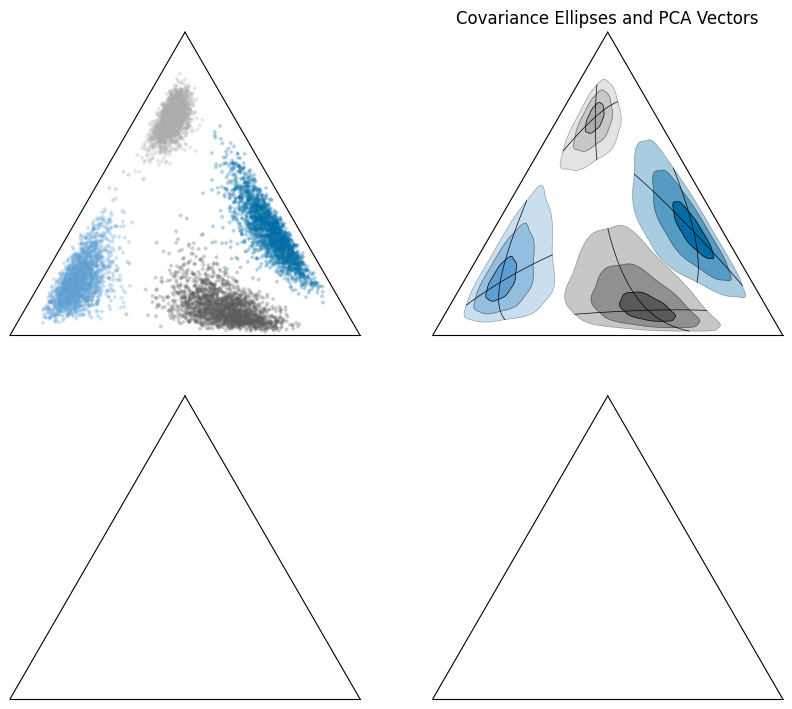
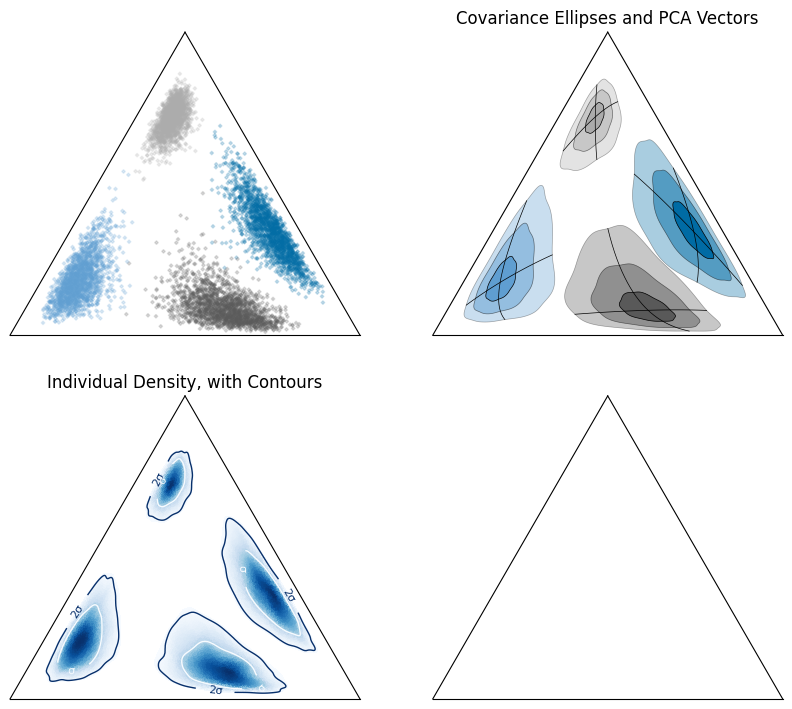
First, let’s look at the synthetic data itself in the ternary space:

We can take the mean and covariance in log-space to create covariance ellipses and vectors using principal component analysis:
kwargs = dict(ax=ax[1], transform=from_log, nstds=3)
ax[1].set_title("Covariance Ellipses and PCA Vectors")
for ix, sample in enumerate(df.Sample.unique()):
comp = df.query("Sample == {}".format(sample))
tcomp = to_log(comp.loc[:, chem])
plot_stdev_ellipses(tcomp.values, color=t10b3[ix], resolution=1000, **kwargs)
plot_pca_vectors(tcomp.values, ls="-", lw=0.5, color="k", **kwargs)
plt.show()

We can also look at data density (here using kernel density estimation) in logratio-space:
kwargs = dict(ax=ax[-2], bins=100, axlabels=False)
ax[-2].set_title("Individual Density, with Contours")
for ix, sample in enumerate(df.Sample.unique()):
comp = df.query("Sample == {}".format(sample))
comp.loc[:, chem].pyroplot.density(cmap="Blues", vmin=0.05, **kwargs)
comp.loc[:, chem].pyroplot.density(
contours=[0.68, 0.95],
cmap="Blues_r",
contour_labels={0.68: "σ", 0.95: "2σ"},
**kwargs,
)
plt.show()
/home/docs/checkouts/readthedocs.org/user_builds/pyrolite/checkouts/develop/pyrolite/comp/codata.py:177: RuntimeWarning: invalid value encountered in log
Y = np.log(X) # Log operation
/home/docs/checkouts/readthedocs.org/user_builds/pyrolite/checkouts/develop/pyrolite/comp/codata.py:177: RuntimeWarning: invalid value encountered in log
Y = np.log(X) # Log operation
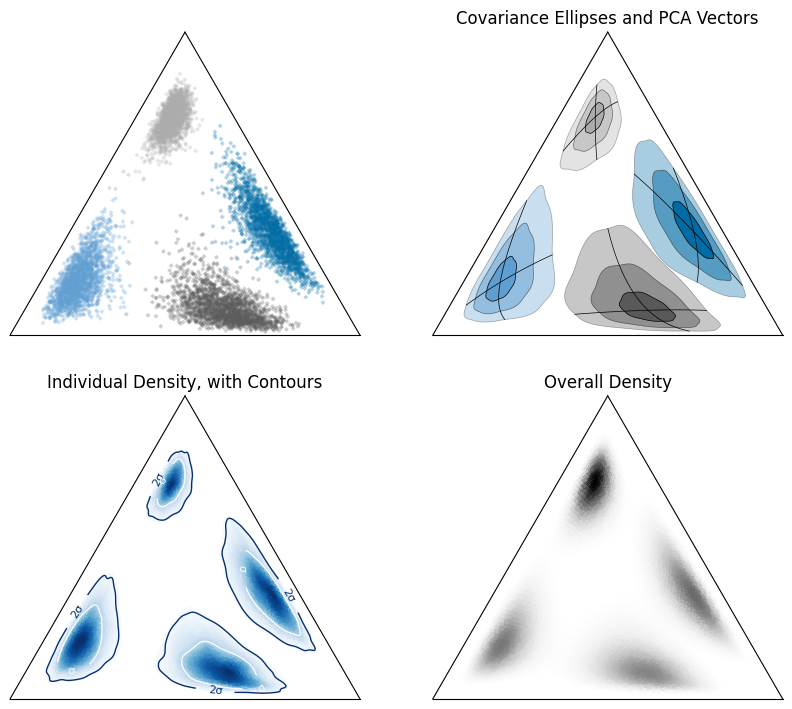
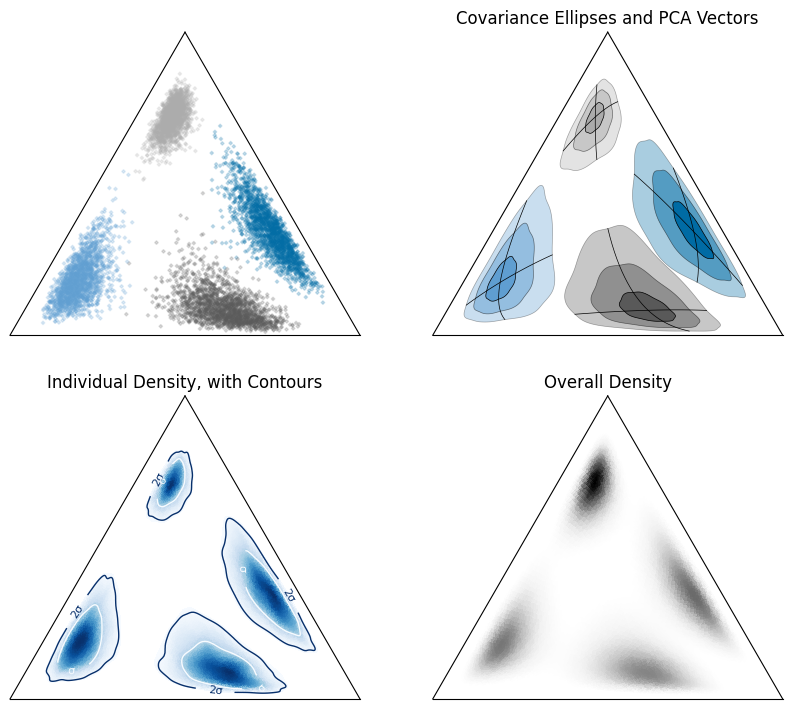
We can also do this for individual samples, and estimate percentile contours:
for a in ax:
a.set_aspect("equal")
a.patch.set_visible(False)
plt.show()
Total running time of the script: (0 minutes 11.255 seconds)